Note
Click here to download the full example code
PPG Pre-processing and SQI Calculations using Pandas¶
Trimming, Filtering, Segmentation and SQI Calculations utilising Pandas
9 10 11 12 | # Code Adapted from:
# https://meta00.github.io/vital_sqi/_examples/others/plot_pipeline_02.html
# AND
# https://meta00.github.io/vital_sqi/_examples/others/plot_read_signal.html#sphx-glr-examples-others-plot-read-signal-py
|
16 17 18 19 20 21 | '''
This file will be used in calculating the SQIs for each patient (and discarding parts of the signal soon).
A loop will be implemented in future iterations for the different patiend files. In general, the file is built
parametrically so that the procedure can be automated.
'''
|
Out:
'\nThis file will be used in calculating the SQIs for each patient (and discarding parts of the signal soon).\nA loop will be implemented in future iterations for the different patiend files. In general, the file is built\nparametrically so that the procedure can be automated.\n\n'
Importing Libraries
25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 | # Generic
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import signal
from datetime import datetime
# Scipy
from scipy.stats import skew, kurtosis, entropy
# vitalSQI
from vital_sqi.data.signal_io import PPG_reader
import vital_sqi.highlevel_functions.highlevel as sqi_hl
import vital_sqi.data.segment_split as sqi_sg
from vital_sqi.common.rpeak_detection import PeakDetector
from vital_sqi.preprocess.band_filter import BandpassFilter
import vital_sqi.sqi as sq
|
Loading data
48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 | # Filepath definition
filepath = r'..\..\..\..\OUCRU\01NVa_Dengue\Adults\01NVa-003-2001\PPG'
filename = r'01NVa-003-2001 Smartcare.csv'
filename_Clinical = r'..\..\..\..\OUCRU\Clinical\v0.0.10\01nva_data_stacked_corrected.csv'
#defining constants
trim_amount = 300
hp_filt_params = (1, 1) #(Hz, order)
lp_filt_params = (20, 4) #(Hz, order)
filter_type = 'butter'
segment_length = 30
sampling_rate = 100
#reading Clinical Data
Clinical = pd.read_csv(filename_Clinical)
#readind PPG data
data = pd.read_csv(os.path.join(filepath, filename))
# data = PPG_reader(os.path.join(filepath, filename),
# signal_idx=["PLETH", "IR_ADC"],
# timestamp_idx=["TIMESTAMP_MS"],
# info_idx=["SPO2_PCT","PULSE_BPM","PERFUSION_INDEX"], sampling_rate=sampling_rate)
print(data)
print(data.PLETH)
print(data.IR_ADC)
print(data.TIMESTAMP_MS)
print(data.SPO2_PCT)
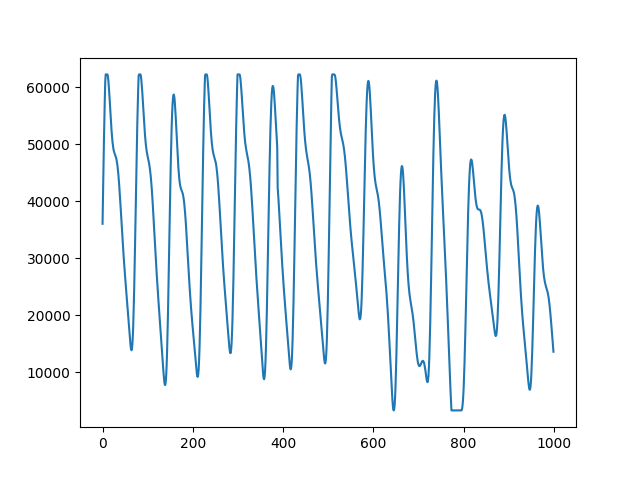
signals = data[["PLETH", "IR_ADC"]]
plot_range = np.arange(0,1000,1)
fig, ax = plt.subplots()
ax.plot(plot_range, signals.iloc[0:1000, 0])
|
Out:
TIMESTAMP_MS COUNTER DEVICE_ID PULSE_BPM SPO2_PCT SPO2_STATUS PLETH BATTERY_PCT RED_ADC IR_ADC PERFUSION_INDEX
0 0 9 1 82 97.1 0 36033 100.0 144180 216524 4.4
1 10 10 1 82 97.1 0 40919 100.0 144160 216470 4.4
2 20 11 1 82 97.1 0 45715 100.0 144158 216415 4.4
3 30 12 1 82 97.1 0 50227 100.0 144148 216364 4.4
4 40 13 1 82 97.1 0 54362 100.0 144132 216379 4.4
... ... ... ... ... ... ... ... ... ... ... ...
5626372 56269900 5626999 1 78 91.4 0 31408 5.7 292213 183755 5.6
5626373 56269910 5627000 1 78 91.4 0 31318 5.7 292557 184002 5.6
5626374 56269920 5627001 1 78 91.4 0 31117 5.7 292840 184296 5.6
5626375 56269930 5627002 1 78 91.4 0 30761 5.7 293161 184554 5.6
5626376 56269940 5627003 1 78 91.4 0 30261 5.7 293522 184828 5.6
[5626377 rows x 11 columns]
0 36033
1 40919
2 45715
3 50227
4 54362
...
5626372 31408
5626373 31318
5626374 31117
5626375 30761
5626376 30261
Name: PLETH, Length: 5626377, dtype: int64
0 216524
1 216470
2 216415
3 216364
4 216379
...
5626372 183755
5626373 184002
5626374 184296
5626375 184554
5626376 184828
Name: IR_ADC, Length: 5626377, dtype: int64
0 0
1 10
2 20
3 30
4 40
...
5626372 56269900
5626373 56269910
5626374 56269920
5626375 56269930
5626376 56269940
Name: TIMESTAMP_MS, Length: 5626377, dtype: int64
0 97.1
1 97.1
2 97.1
3 97.1
4 97.1
...
5626372 91.4
5626373 91.4
5626374 91.4
5626375 91.4
5626376 91.4
Name: SPO2_PCT, Length: 5626377, dtype: float64
[<matplotlib.lines.Line2D object at 0x000001C5BCEA4B50>]
fetching ppg_start time
91 92 93 | def find_event_ppg_time(Clinical,Patient):
row = Clinical[(Clinical.study_no == Patient) & (Clinical.column == 'event_ppg') & (Clinical.result == 'True')]
return row['date'].values
|
Setting the indexes for the raw data - adopted from examples
97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 | pd.Timedelta.__str__ = lambda x: x._repr_base('all')
#Transposing data.signals to set into columns
#signals = pd.DataFrame(data.signals.T)
# Include column with index
signals = signals.reset_index()
signals['timedelta'] = pd.to_timedelta(data.TIMESTAMP_MS, unit='ms')
PPG_start_date = find_event_ppg_time(Clinical, filename.split()[0][-8:])
PPG_start_date = datetime.strptime(PPG_start_date[0], '%Y-%m-%d %H:%M:%S')
signals['PPG_Datetime'] = pd.to_datetime(PPG_start_date)
signals['PPG_Datetime']+= pd.to_timedelta(signals.timedelta)
# Set the timedelta index (keep numeric index too)
signals = signals.set_index('timedelta')
# Rename column to avoid confusion
signals = signals.rename(columns={'index': 'idx'})
print("Raw Signals:")
print(signals)
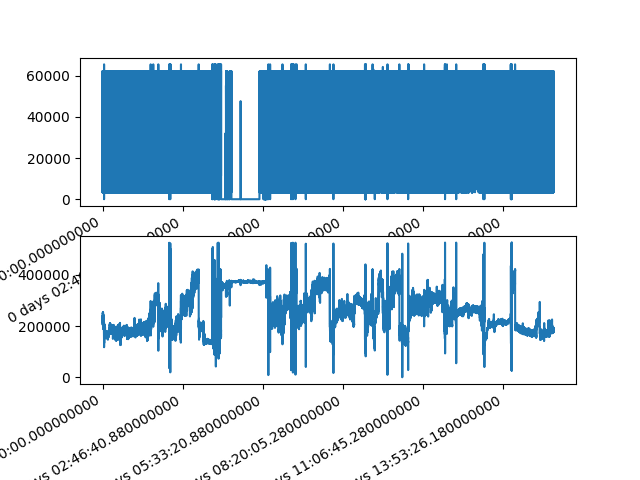
#Plotting
fig, axes = plt.subplots(nrows=2, ncols=1)
axes = axes.flatten()
signals.iloc[:,1].plot(ax=axes[0])
signals.iloc[:,2].plot(ax=axes[1])
|
Out:
Raw Signals:
idx PLETH IR_ADC PPG_Datetime
timedelta
0 days 00:00:00 0 36033 216524 2020-06-04 09:05:00.000
0 days 00:00:00.010000 1 40919 216470 2020-06-04 09:05:00.010
0 days 00:00:00.020000 2 45715 216415 2020-06-04 09:05:00.020
0 days 00:00:00.030000 3 50227 216364 2020-06-04 09:05:00.030
0 days 00:00:00.040000 4 54362 216379 2020-06-04 09:05:00.040
... ... ... ... ...
0 days 15:37:49.900000 5626372 31408 183755 2020-06-05 00:42:49.900
0 days 15:37:49.910000 5626373 31318 184002 2020-06-05 00:42:49.910
0 days 15:37:49.920000 5626374 31117 184296 2020-06-05 00:42:49.920
0 days 15:37:49.930000 5626375 30761 184554 2020-06-05 00:42:49.930
0 days 15:37:49.940000 5626376 30261 184828 2020-06-05 00:42:49.940
[5626377 rows x 4 columns]
<AxesSubplot:xlabel='timedelta'>
Trimming the first and last 5 minutes
132 133 134 135 136 137 138 139 140 141 142 | #Defining the 5 minute offset
offset = pd.Timedelta(minutes=5)
#Indexes - adopted from example
idxs = (signals.index > offset) & \
(signals.index < signals.index[-1] - offset)
#Trimming
signals = signals[idxs]
print(signals)
|
Out:
idx PLETH IR_ADC PPG_Datetime
timedelta
0 days 00:05:00.010000 29913 21755 188656 2020-06-04 09:10:00.010
0 days 00:05:00.020000 29914 23262 187701 2020-06-04 09:10:00.020
0 days 00:05:00.030000 29915 25285 186998 2020-06-04 09:10:00.030
0 days 00:05:00.040000 29916 27839 186472 2020-06-04 09:10:00.040
0 days 00:05:00.050000 29917 30888 186113 2020-06-04 09:10:00.050
... ... ... ... ...
0 days 15:32:49.890000 5596371 15083 186186 2020-06-05 00:37:49.890
0 days 15:32:49.900000 5596372 13923 186422 2020-06-05 00:37:49.900
0 days 15:32:49.910000 5596373 12818 186662 2020-06-05 00:37:49.910
0 days 15:32:49.920000 5596374 11769 186807 2020-06-05 00:37:49.920
0 days 15:32:49.930000 5596375 10763 186899 2020-06-05 00:37:49.930
[5566463 rows x 4 columns]
147 148 149 | # RESAMPLE??
# MISSING DATA INPUT??
# TAPPERING??
|
Band-pass filtering
154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 | '''
Band-pass filtering can be adjusted, maybe use a lower low-pass
cutoff frequency @15-17Hz
'''
#Band pass filtering function
def bpf(signal):
hp_filt_params = (1, 1) #(Cutoff Hz, order)
lp_filt_params = (20, 4) #( Cutoff Hz, order)
sampling_rate = 100
filter = BandpassFilter(band_type='butter', fs=sampling_rate)
filtered_signal = filter.signal_highpass_filter(signal, cutoff=hp_filt_params[0], order=hp_filt_params[1])
filtered_signal = filter.signal_lowpass_filter(filtered_signal, cutoff=lp_filt_params[0], order=lp_filt_params[1])
return filtered_signal
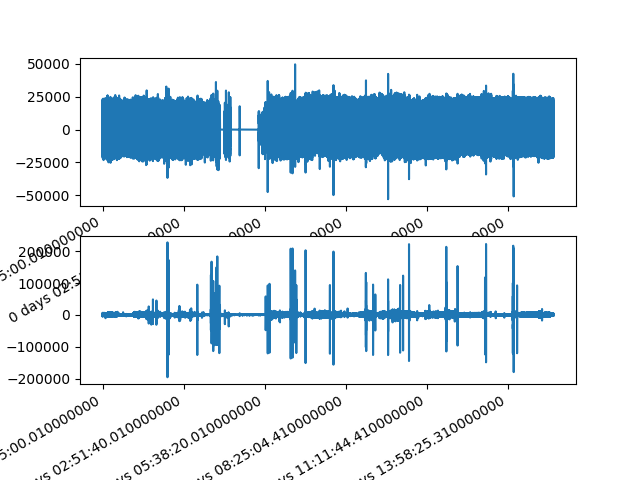
#Band-pass filtered signals
signals['PLETH_bpf'] = bpf(signals.iloc[:,1])
signals['IR_ADC_bpf'] = bpf(signals.iloc[:,2])
print('Raw and Filtered Signals:')
print(signals)
fig, axes = plt.subplots(nrows=2, ncols=1)
axes = axes.flatten()
signals['PLETH_bpf'].plot(ax=axes[0])
signals['IR_ADC_bpf'].plot(ax=axes[1])
|
Out:
Raw and Filtered Signals:
idx PLETH IR_ADC PPG_Datetime PLETH_bpf IR_ADC_bpf
timedelta
0 days 00:05:00.010000 29913 21755 188656 2020-06-04 09:10:00.010 -214.886697 -32.494545
0 days 00:05:00.020000 29914 23262 187701 2020-06-04 09:10:00.020 -1224.999347 -226.831554
0 days 00:05:00.030000 29915 25285 186998 2020-06-04 09:10:00.030 -3047.445501 -734.950280
0 days 00:05:00.040000 29916 27839 186472 2020-06-04 09:10:00.040 -4351.721824 -1499.309485
0 days 00:05:00.050000 29917 30888 186113 2020-06-04 09:10:00.050 -3863.561589 -2230.015638
... ... ... ... ... ... ...
0 days 15:32:49.890000 5596371 15083 186186 2020-06-05 00:37:49.890 942.030869 45.325519
0 days 15:32:49.900000 5596372 13923 186422 2020-06-05 00:37:49.900 708.195771 93.296871
0 days 15:32:49.910000 5596373 12818 186662 2020-06-05 00:37:49.910 542.754581 127.077955
0 days 15:32:49.920000 5596374 11769 186807 2020-06-05 00:37:49.920 442.131641 151.598399
0 days 15:32:49.930000 5596375 10763 186899 2020-06-05 00:37:49.930 402.889073 159.679506
[5566463 rows x 6 columns]
<AxesSubplot:xlabel='timedelta'>
SQI Functions Definition and Peak Detection
185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 | def PeakDetection(x):
detector = PeakDetector()
peaks, troughs = detector.ppg_detector(x, 7) #6 and 7 work the best
return peaks, troughs
#Defining SQI Functions
def snr(x):
#Signal to noise ratio
#needs raw signal
return np.mean(sq.standard_sqi.signal_to_noise_sqi(x))
def zcr(x):
#Zero Crossing Rate
#Needs filtered signal
return 0.5 * np.mean(np.abs(np.diff(np.sign(x))))
def mcr(x):
#Mean Crossing Rate
#needs raw signal
return zcr(x - np.mean(x))
def perfusion(x,y):
return (np.max(y) - np.min(y)) / np.abs(np.mean(x)) * 100
def correlogram(x):
#Correlogram
#Needs filtered signal
return sq.rpeaks_sqi.correlogram_sqi(x)
#Calculating peaks and troughs using peakdetector function for both filtered signals
peak_list_0, trough_list_0 = PeakDetection(signals['PLETH_bpf'])
peak_list_1, trough_list_1 = PeakDetection(signals['IR_ADC_bpf'])
'''
MSW values that are being calculated are too small, maybe peakdetectors need
to be revisited?
'''
def msq(x,peaks):
#Dynamic Time Warping
#Requires Filtered Signal and peaks
# Return
return sq.standard_sqi.msq_sqi(x, peaks_1=peaks, peak_detect2=6)
'''
Below function is outputting an error, needs revisiting
'''
# def dtw(x,troughs):
# #Dynamic Time Warping
# #Requires Filtered Signal and troughs and template selection 0-3 (0-2 for PPG)
# # Per beat
# dtw_list = sq.standard_sqi.per_beat_sqi(sqi_func=sq.dtw_sqi, troughs=troughs, signal=x, taper=True, template_type=1)
# # Return mean and standard deviation
# return [np.mean(dtw_list), np.std(dtw_list)]
|
Out:
'\nBelow function is outputting an error, needs revisiting\n'
Function Computing SQIs
243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 | def sqi_all(x,raw,filtered,peaks,troughs):
#Computing all SQIs using the functions above
# Information
dinfo = {
'PPG_w_s': x.PPG_Datetime.iloc[0],
'PPG_w_f': x.PPG_Datetime.iloc[-1],
'first': x.idx.iloc[0],
'last': x.idx.iloc[-1],
'skew': skew(x[raw]),
'entropy': entropy(x[raw]),
'kurtosis': kurtosis(x[raw]),
'snr': snr(x[raw]),
'mcr': mcr(x[raw]),
'zcr': zcr(x[filtered]),
'msq': msq(x[filtered],peaks),
'perfusion': perfusion(x[raw], x[filtered]),
'correlogram': correlogram(x[filtered]),
#'dtw': dtw(x[filtered],troughs)
}
'''
When using dtw an empty signal error is presented that
needs fixing, not sure why it does that
'''
# Return
return pd.Series(dinfo)
|
Calculating SQIs for both signals 0 = Pleth and 1 = IR_ADC
274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 | sqis_0 = signals.groupby(pd.Grouper(freq='30s')).apply(lambda x: sqi_all(x, 'PLETH','PLETH_bpf', peak_list_0, trough_list_0))
sqis_1 = signals.groupby(pd.Grouper(freq='30s')).apply(lambda x: sqi_all(x, 'IR_ADC','IR_ADC_bpf', peak_list_1, trough_list_1))
#Displaying the SQIs per signal
print("SQIs Signal 0 (Pleth):")
sqis_0
print(sqis_0)
print("SQIs Signal 1 (IR_ADC):")
sqis_1
print(sqis_1)
sqis_1 = sqis_1.drop(['first','last', 'PPG_w_s', 'PPG_w_f'], axis = 1)
# Add window id to identify point of shock more easily
sqis_1['w'] = np.arange(sqis_1.shape[0])
|
Out:
d:\files\desktop\dissertation icl\env\lib\site-packages\statsmodels\tsa\stattools.py:667: FutureWarning: fft=True will become the default after the release of the 0.12 release of statsmodels. To suppress this warning, explicitly set fft=False.
warnings.warn(
SQIs Signal 0 (Pleth):
PPG_w_s PPG_w_f first ... msq perfusion correlogram
timedelta ...
0 days 00:05:00.010000 2020-06-04 09:10:00.010 2020-06-04 09:10:30.000 29913 ... 0.000438 131.768567 [75, 150, 225, 0.9247323846304137, 0.836659565...
0 days 00:05:30.010000 2020-06-04 09:10:30.010 2020-06-04 09:11:00.000 32913 ... 0.000037 129.721286 [74, 148, 222, 0.9374430072923502, 0.866928327...
0 days 00:06:00.010000 2020-06-04 09:11:00.010 2020-06-04 09:11:30.000 35913 ... 0.000000 135.690506 [75, 150, 225, 0.9244827555340438, 0.814277364...
0 days 00:06:30.010000 2020-06-04 09:11:30.010 2020-06-04 09:12:00.000 38913 ... 0.000007 136.156223 [75, 150, 225, 0.9386014677175014, 0.862369440...
0 days 00:07:00.010000 2020-06-04 09:12:00.010 2020-06-04 09:12:30.000 41913 ... 0.000000 139.046179 [74, 149, 224, 0.9313743004418997, 0.824739899...
... ... ... ... ... ... ... ...
0 days 15:30:30.010000 2020-06-05 00:35:30.010 2020-06-05 00:36:00.000 5582383 ... 0.000000 148.508917 [37, 74, 110, -0.3912006306694429, 0.952924647...
0 days 15:31:00.010000 2020-06-05 00:36:00.010 2020-06-05 00:36:30.000 5585383 ... 0.000022 138.961305 [37, 75, 112, -0.38664616039771205, 0.95837555...
0 days 15:31:30.010000 2020-06-05 00:36:30.010 2020-06-05 00:37:00.000 5588383 ... 0.000015 125.616461 [77, 153, 230, 0.9462808953640326, 0.887181765...
0 days 15:32:00.010000 2020-06-05 00:37:00.010 2020-06-05 00:37:30.000 5591383 ... 0.000015 135.269077 [36, 74, 110, -0.37596691904957935, 0.93875374...
0 days 15:32:30.010000 2020-06-05 00:37:30.010 2020-06-05 00:37:49.930 5594383 ... 0.000022 134.870996 [37, 75, 112, -0.36633265827700123, 0.94094953...
[1856 rows x 13 columns]
SQIs Signal 1 (IR_ADC):
PPG_w_s PPG_w_f first ... msq perfusion correlogram
timedelta ...
0 days 00:05:00.010000 2020-06-04 09:10:00.010 2020-06-04 09:10:30.000 29913 ... 0.000319 6.653209 [75, 150, 226, 0.9132608947040779, 0.799647163...
0 days 00:05:30.010000 2020-06-04 09:10:30.010 2020-06-04 09:11:00.000 32913 ... 0.000013 5.641682 [74, 148, 222, 0.9164960001123641, 0.827303342...
0 days 00:06:00.010000 2020-06-04 09:11:00.010 2020-06-04 09:11:30.000 35913 ... 0.000008 6.088179 [75, 150, 225, 0.9097936306566979, 0.779681593...
0 days 00:06:30.010000 2020-06-04 09:11:30.010 2020-06-04 09:12:00.000 38913 ... 0.000013 4.644686 [75, 149, 224, 0.9128541336509863, 0.807913379...
0 days 00:07:00.010000 2020-06-04 09:12:00.010 2020-06-04 09:12:30.000 41913 ... 0.000000 4.241945 [74, 149, 223, 0.9123059092032415, 0.791340070...
... ... ... ... ... ... ... ...
0 days 15:30:30.010000 2020-06-05 00:35:30.010 2020-06-05 00:36:00.000 5582383 ... 0.000013 4.946963 [34, 74, 107, -0.38922458037787383, 0.95155578...
0 days 15:31:00.010000 2020-06-05 00:36:00.010 2020-06-05 00:36:30.000 5585383 ... 0.000000 4.268080 [75, 150, 226, 0.964216933015672, 0.9155762504...
0 days 15:31:30.010000 2020-06-05 00:36:30.010 2020-06-05 00:37:00.000 5588383 ... 0.000013 4.733346 [77, 153, 230, 0.9423859574341878, 0.877431048...
0 days 15:32:00.010000 2020-06-05 00:37:00.010 2020-06-05 00:37:30.000 5591383 ... 0.000008 4.645047 [74, 103, 148, 0.9269084193473138, -0.35288054...
0 days 15:32:30.010000 2020-06-05 00:37:30.010 2020-06-05 00:37:49.930 5594383 ... 0.000008 4.352384 [75, 150, 226, 0.9326593121930683, 0.859995140...
[1856 rows x 13 columns]
Final Formatting into a single DataFrame and Saving into a .csv file
295 296 297 298 299 300 301 302 303 | #Merging SQIs of different signals from the same record (Pleth and IR_ADC)
Signal_SQIs = sqis_0.merge(sqis_1, on='timedelta', suffixes=('_0', '_1'))
#Automatically fetching the study_no from the filename and including it in the DataFrame
Signal_SQIs['study_no'] = filename.split()[0][-8:]
print(Signal_SQIs)
# Saving the SQIs to a CSV file
Signal_SQIs.to_csv(r'..\..\..\..\OUCRU\Outputs\SQI.csv')
|
Out:
PPG_w_s PPG_w_f first ... correlogram_1 w study_no
timedelta ...
0 days 00:05:00.010000 2020-06-04 09:10:00.010 2020-06-04 09:10:30.000 29913 ... [75, 150, 226, 0.9132608947040779, 0.799647163... 0 003-2001
0 days 00:05:30.010000 2020-06-04 09:10:30.010 2020-06-04 09:11:00.000 32913 ... [74, 148, 222, 0.9164960001123641, 0.827303342... 1 003-2001
0 days 00:06:00.010000 2020-06-04 09:11:00.010 2020-06-04 09:11:30.000 35913 ... [75, 150, 225, 0.9097936306566979, 0.779681593... 2 003-2001
0 days 00:06:30.010000 2020-06-04 09:11:30.010 2020-06-04 09:12:00.000 38913 ... [75, 149, 224, 0.9128541336509863, 0.807913379... 3 003-2001
0 days 00:07:00.010000 2020-06-04 09:12:00.010 2020-06-04 09:12:30.000 41913 ... [74, 149, 223, 0.9123059092032415, 0.791340070... 4 003-2001
... ... ... ... ... ... ... ...
0 days 15:30:30.010000 2020-06-05 00:35:30.010 2020-06-05 00:36:00.000 5582383 ... [34, 74, 107, -0.38922458037787383, 0.95155578... 1851 003-2001
0 days 15:31:00.010000 2020-06-05 00:36:00.010 2020-06-05 00:36:30.000 5585383 ... [75, 150, 226, 0.964216933015672, 0.9155762504... 1852 003-2001
0 days 15:31:30.010000 2020-06-05 00:36:30.010 2020-06-05 00:37:00.000 5588383 ... [77, 153, 230, 0.9423859574341878, 0.877431048... 1853 003-2001
0 days 15:32:00.010000 2020-06-05 00:37:00.010 2020-06-05 00:37:30.000 5591383 ... [74, 103, 148, 0.9269084193473138, -0.35288054... 1854 003-2001
0 days 15:32:30.010000 2020-06-05 00:37:30.010 2020-06-05 00:37:49.930 5594383 ... [75, 150, 226, 0.9326593121930683, 0.859995140... 1855 003-2001
[1856 rows x 24 columns]
308 309 310 311 312 313 | '''
#Running Some Abstract Checks to Verify Results etc.
print(perfusion(signals[0][0:3000], signals['PLETH_bpf'][0:3000]))
'''
|
Out:
"\n#Running Some Abstract Checks to Verify Results etc.\n\nprint(perfusion(signals[0][0:3000], signals['PLETH_bpf'][0:3000]))\n\n"
316 317 318 | '''
Discarding Process to be implemented after quality check.
'''
|
Out:
'\nDiscarding Process to be implemented after quality check.\n'
Total running time of the script: ( 1 minutes 59.605 seconds)