Note
Click here to download the full example code
Matching SQI Dataframe Window to Clinical Time of “Event” for Multiple Records¶
Matching SQI Dataframe Window to Clinical Time of “Event” for Multiple Records
Importing Libraries
12 13 14 15 16 17 | from datetime import date
import pandas as pd
import numpy as np
from datetime import datetime
import matplotlib.pyplot as plt
import os
|
Loading SQI Data and clinical data
22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 | filename_SQIs = r'..\..\..\..\OUCRU\Outputs\Complete_SQIs.csv'
filename_Clinical = r'..\..\..\..\OUCRU\Clinical\v0.0.10\01nva_data_stacked_corrected.csv'
study_no_path = r'..\..\..\..\OUCRU\01NVa_Dengue\Adults'
#taking a list of records
study_no_list = os.listdir(study_no_path) # Need to take the last 8 characters i.e. study_no_list[i][-8:]
#reading .csv into dataframes
Clinical = pd.read_csv(filename_Clinical)
SQIs = pd.read_csv(filename_SQIs)
SQIs['PPG_w_s'] = pd.to_datetime(SQIs['PPG_w_s'])
SQIs['PPG_w_f'] = pd.to_datetime(SQIs['PPG_w_f'])
TERMINAL = True
#Showing Data
if TERMINAL:
print("\n Clincial Data:")
print(Clinical)
print("\n SQI Data:")
print(SQIs)
def match_clinical_to_SQIs(Clinical, SQIs, event, study_no_list):
SQIs[event]=np.nan #forming an empty column for the event
print("\n EVENT: ", event)
#iterating through the study_no
for i in range(len(study_no_list)):
print("\n STUDY-NO: ", study_no_list[i])
#finding the rows for which the following logic is satisfied
event_row = Clinical[(((Clinical.column == event) & (Clinical.result == 'TRUE')) | (Clinical.result == event)) & (Clinical.study_no == study_no_list[i][-8:])]
#iterating through the event row for indexing
event_row['date'] = pd.to_datetime(event_row['date'])
if event == 'shock_admission' and not event_row.empty:
SQIs['shock_admission'][SQIs.study_no == study_no_list[i][-8:]] = True #Showing whether a record indicates a shock on admission
else:
for j in range(len(event_row)):
valid_SQI_rows = SQIs[(SQIs.PPG_w_s <= event_row.date[event_row.date.index[j]]) & (SQIs.PPG_w_f > event_row.date[event_row.date.index[j]]) & (SQIs.study_no == study_no_list[i][-8:])]
SQIs[event][valid_SQI_rows.index] = True #Setting the value to True for the window corresponding to an event
if valid_SQI_rows.empty:
##If the date is equal to the date of an event, we can raise True for the whole day, in case no time is present (or it's out of the recording)
SQIs[event][(SQIs.PPG_w_s.dt.date == event_row.date[event_row.date.index[j]].date) & (SQIs.PPG_w_f.dt.date == event_row.date[event_row.date.index[j]].date) & (SQIs.study_no == study_no_list[i][-8:])]
return SQIs
'''
NOTE:
We previously had functions that fetched ppg start and
calculated relative times but since we included the PPG Start times
in the SQI file, that is no longer needed.
'''
#Calling the matching function by first defining the event
event_lookup = 'event_shock'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs, event_lookup, study_no_list)
event_lookup = 'reshock24'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'diagnosis_admission' #all diagnosis admission are for shock so we can use this as is
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'shock_admission'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
#In the second iteration we call the function with the output of the first one, so that we can form
#multiple columns of events
event_lookup = 'ascites'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'respiratory_distress'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'ventilation_cannula'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'ventilation_mechanical'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'ventilation_ncpap'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'bleeding_severe'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'cns_abnormal'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'liver_mild'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'pleural_effusion'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
event_lookup = 'skidney'
SQIs_with_Clinical = match_clinical_to_SQIs(Clinical, SQIs_with_Clinical, event_lookup, study_no_list)
if TERMINAL:
print('\n SQIs with Clinical Event Match:')
print(SQIs_with_Clinical)
#Saving output to CSV
SQIs_with_Clinical.to_csv(r'..\..\..\..\OUCRU\Outputs\Complete_SQIs_with_Clinical.csv')
if TERMINAL:
#example of the output
fig, axs = plt.subplots(nrows=2, sharex=True, figsize=(8, 10))
img = plt.imread(r'..\..\..\..\MISC\SQI_Clinical_Image_example_2.png')
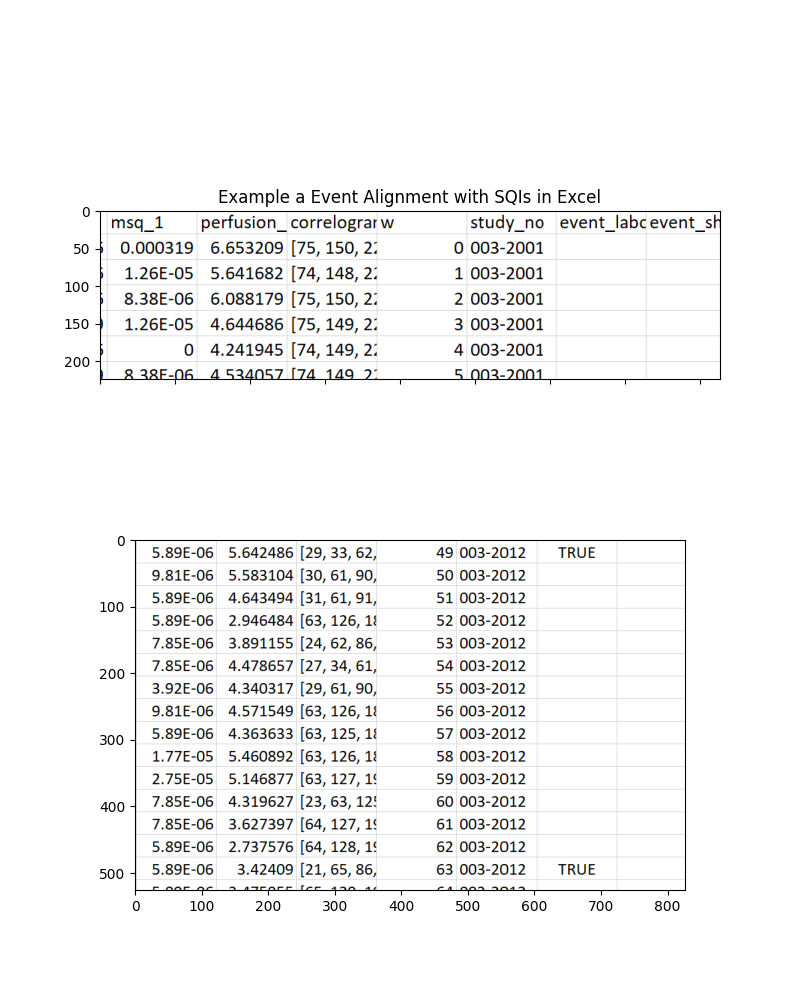
axs[0].set_title('Example a Event Alignment with SQIs in Excel')
axs[0].imshow(img)
img2 = plt.imread(r'..\..\..\..\MISC\SQI_Clinical_Image_example_3.png')
axs[1].imshow(img2)
plt.show()
|
Out:
Clincial Data:
Unnamed: 0 study_no date column result unit result_old date_old
0 0 003-1001 11/06/2020 13:30 hypertension TRUE NaN NaN NaN
1 1 003-1002 17/06/2020 11:10 hypertension FALSE NaN NaN NaN
2 2 003-1003 18/06/2020 07:30 hypertension FALSE NaN NaN NaN
3 3 003-1004 22/06/2020 12:45 hypertension FALSE NaN NaN NaN
4 4 003-1005 02/07/2020 15:11 hypertension FALSE NaN NaN NaN
... ... ... ... ... ... ... ... ...
42060 42060 003-2231 05/12/2020 00:00 outcome Full recovery NaN NaN NaN
42061 42061 003-2232 19/12/2020 00:00 outcome Full recovery NaN NaN NaN
42062 42062 003-2233 02/01/2021 00:00 outcome Full recovery NaN NaN NaN
42063 42063 003-2234 08/01/2021 00:00 outcome Full recovery NaN NaN NaN
42064 42064 003-2235 18/01/2021 00:00 outcome Full recovery NaN NaN NaN
[42065 rows x 8 columns]
SQI Data:
timedelta PPG_w_s PPG_w_f first ... w study_no study_no_rec keep
0 0 days 00:05:00.010000 2020-07-28 16:04:20.104 2020-07-28 16:04:50.094 29913 ... 0 003-2009 0 True
1 0 days 00:05:30.010000 2020-07-28 16:04:50.104 2020-07-28 16:05:20.094 32913 ... 1 003-2009 0 True
2 0 days 00:06:00.010000 2020-07-28 16:05:20.104 2020-07-28 16:05:50.094 35913 ... 2 003-2009 0 True
3 0 days 00:06:30.010000 2020-07-28 16:05:50.104 2020-07-28 16:06:20.094 38913 ... 3 003-2009 0 True
4 0 days 00:07:00.010000 2020-07-28 16:06:20.104 2020-07-28 16:06:50.094 41913 ... 4 003-2009 0 True
... ... ... ... ... ... ... ... ... ...
12698 0 days 15:03:30.010000 2020-07-21 05:08:37.203 2020-07-21 05:09:07.193 5420828 ... 1797 003-2162 0 True
12699 0 days 15:04:00.010000 2020-07-21 05:09:07.203 2020-07-21 05:09:37.193 5423828 ... 1798 003-2162 0 True
12700 0 days 15:04:30.010000 2020-07-21 05:09:37.203 2020-07-21 05:10:07.193 5426828 ... 1799 003-2162 0 True
12701 0 days 15:05:00.010000 2020-07-21 05:10:07.203 2020-07-21 05:10:37.193 5429828 ... 1800 003-2162 0 True
12702 0 days 15:05:30.010000 2020-07-21 05:10:37.203 2020-07-21 05:10:52.883 5432828 ... 1801 003-2162 0 True
[12703 rows x 27 columns]
EVENT: event_shock
STUDY-NO: 01NVa-003-2009
D:\FILES\Desktop\Dissertation ICL\Git\main\examples\Pre-processing\plot_SQI_and_clinical_matching_multiple_patients.py:56: SettingWithCopyWarning:
A value is trying to be set on a copy of a slice from a DataFrame.
Try using .loc[row_indexer,col_indexer] = value instead
See the caveats in the documentation: https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy
D:\FILES\Desktop\Dissertation ICL\Git\main\examples\Pre-processing\plot_SQI_and_clinical_matching_multiple_patients.py:62: SettingWithCopyWarning:
A value is trying to be set on a copy of a slice from a DataFrame
See the caveats in the documentation: https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: reshock24
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: diagnosis_admission
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: shock_admission
STUDY-NO: 01NVa-003-2009
D:\FILES\Desktop\Dissertation ICL\Git\main\examples\Pre-processing\plot_SQI_and_clinical_matching_multiple_patients.py:58: SettingWithCopyWarning:
A value is trying to be set on a copy of a slice from a DataFrame
See the caveats in the documentation: https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: ascites
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: respiratory_distress
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: ventilation_cannula
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: ventilation_mechanical
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: ventilation_ncpap
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: bleeding_severe
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: cns_abnormal
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: liver_mild
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: pleural_effusion
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
EVENT: skidney
STUDY-NO: 01NVa-003-2009
STUDY-NO: 01NVa-003-2012
STUDY-NO: 01NVa-003-2023
STUDY-NO: 01NVa-003-2028
STUDY-NO: 01NVa-003-2103
STUDY-NO: 01NVa-003-2104
STUDY-NO: 01NVa-003-2109
STUDY-NO: 01NVa-003-2110
STUDY-NO: 01NVa-003-2162
SQIs with Clinical Event Match:
timedelta PPG_w_s PPG_w_f first ... cns_abnormal liver_mild pleural_effusion skidney
0 0 days 00:05:00.010000 2020-07-28 16:04:20.104 2020-07-28 16:04:50.094 29913 ... NaN NaN NaN NaN
1 0 days 00:05:30.010000 2020-07-28 16:04:50.104 2020-07-28 16:05:20.094 32913 ... NaN NaN NaN NaN
2 0 days 00:06:00.010000 2020-07-28 16:05:20.104 2020-07-28 16:05:50.094 35913 ... NaN NaN NaN NaN
3 0 days 00:06:30.010000 2020-07-28 16:05:50.104 2020-07-28 16:06:20.094 38913 ... NaN NaN NaN NaN
4 0 days 00:07:00.010000 2020-07-28 16:06:20.104 2020-07-28 16:06:50.094 41913 ... NaN NaN NaN NaN
... ... ... ... ... ... ... ... ... ...
12698 0 days 15:03:30.010000 2020-07-21 05:08:37.203 2020-07-21 05:09:07.193 5420828 ... NaN NaN NaN NaN
12699 0 days 15:04:00.010000 2020-07-21 05:09:07.203 2020-07-21 05:09:37.193 5423828 ... NaN NaN NaN NaN
12700 0 days 15:04:30.010000 2020-07-21 05:09:37.203 2020-07-21 05:10:07.193 5426828 ... NaN NaN NaN NaN
12701 0 days 15:05:00.010000 2020-07-21 05:10:07.203 2020-07-21 05:10:37.193 5429828 ... NaN NaN NaN NaN
12702 0 days 15:05:30.010000 2020-07-21 05:10:37.203 2020-07-21 05:10:52.883 5432828 ... NaN NaN NaN NaN
[12703 rows x 41 columns]
Total running time of the script: ( 0 minutes 4.307 seconds)