Note
Click here to download the full example code
PPG Peak Detector Comparisons¶
Comparing Different Peak Detectors
9 10 11 12 | # Code Adapted from:
# https://github.com/meta00/vital_sqi/blob/main/examples/SQI_pipeline_PPG.ipynb
# AND
# https://meta00.github.io/vital_sqi/_examples/others/plot_read_signal.html#sphx-glr-examples-others-plot-read-signal-py
|
Libraries
17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 | # Generic
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Scipy
from scipy.stats import skew
from scipy.stats import kurtosis
# vitalSQI
from vital_sqi.data.signal_io import PPG_reader
import vital_sqi.highlevel_functions.highlevel as sqi_hl
import vital_sqi.data.segment_split as sqi_sg
from vital_sqi.common.rpeak_detection import PeakDetector
from vital_sqi.preprocess.band_filter import BandpassFilter
import vital_sqi.sqi as sq
|
Out:
Importing the dtw module. When using in academic works please cite:
T. Giorgino. Computing and Visualizing Dynamic Time Warping Alignments in R: The dtw Package.
J. Stat. Soft., doi:10.18637/jss.v031.i07.
Loading data
39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 | # Filepath
filepath = r'..\..\..\..\OUCRU\01NVa_Dengue\Adults\01NVa-003-2001\PPG'
filename = r'01NVa-003-2001 Smartcare.csv'
#defining constants
trim_amount = 300
hp_filt_params = (1, 1) #(Hz, order)
lp_filt_params = (20, 4) #(Hz, order)
filter_type = 'butter'
segment_length = 30
sampling_rate = 100
#readind PPG data
data = PPG_reader(os.path.join(filepath, filename),
signal_idx=["PLETH", "IR_ADC"],
timestamp_idx=["TIMESTAMP_MS"],
info_idx=["SPO2_PCT","PULSE_BPM","PERFUSION_INDEX"], sampling_rate=sampling_rate)
PeakDetectorNames = ["ADAPTIVE_THRESHOLD",
"COUNT_ORIG_METHOD",
"CLUSTERER_METHOD",
"SLOPE_SUM_METHOD",
"MOVING_AVERAGE_METHOD",
"DEFAULT_SCIPY",
"BILLAUER_METHOD"]
|
68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 | pd.Timedelta.__str__ = lambda x: x._repr_base('all')
signals = pd.DataFrame(data.signals.T)
# Include column with index
signals = signals.reset_index()
signals['timedelta'] = \
pd.to_timedelta(signals.index / sampling_rate, unit='s')
signals = signals.set_index('timedelta')
# Show
print("Raw Signals: ")
print(signals)
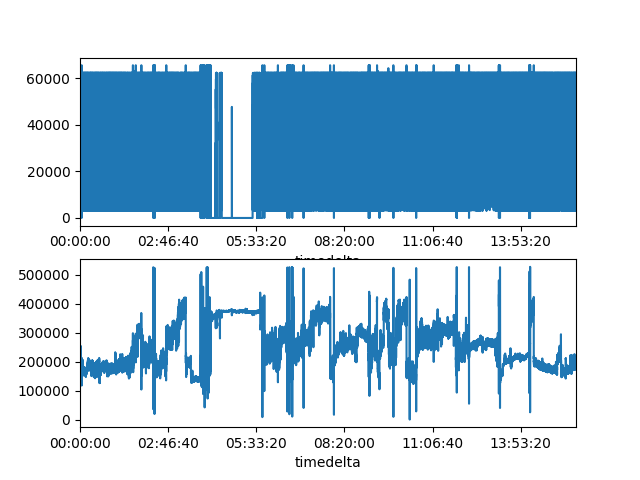
# Plot
fig, axes = plt.subplots(nrows=2, ncols=1)
axes = axes.flatten()
signals[0].plot(ax=axes[0])
signals[1].plot(ax=axes[1])
#==================
#Bandpass Filtering
#==================
def bpf(signal):
hp_filt_params = (1, 1) #(Hz, order)
lp_filt_params = (20, 4) #(Hz, order)
sampling_rate = 100
filter = BandpassFilter(band_type='butter', fs=sampling_rate)
filtered_signal = filter.signal_highpass_filter(signal, cutoff=hp_filt_params[0], order=hp_filt_params[1])
filtered_signal = filter.signal_lowpass_filter(filtered_signal, cutoff=lp_filt_params[0], order=lp_filt_params[1])
return filtered_signal
signals['Filtered_Pleth'] = bpf(signals[0])
signals['Filtered_IR_ADC'] = bpf(signals[1])
|
Out:
Raw Signals:
index 0 1
timedelta
0 days 00:00:00 0 36033 216524
0 days 00:00:00.010000 1 40919 216470
0 days 00:00:00.020000 2 45715 216415
0 days 00:00:00.030000 3 50227 216364
0 days 00:00:00.040000 4 54362 216379
... ... ... ...
0 days 15:37:43.720000 5626372 31408 183755
0 days 15:37:43.730000 5626373 31318 184002
0 days 15:37:43.740000 5626374 31117 184296
0 days 15:37:43.750000 5626375 30761 184554
0 days 15:37:43.760000 5626376 30261 184828
[5626377 rows x 3 columns]
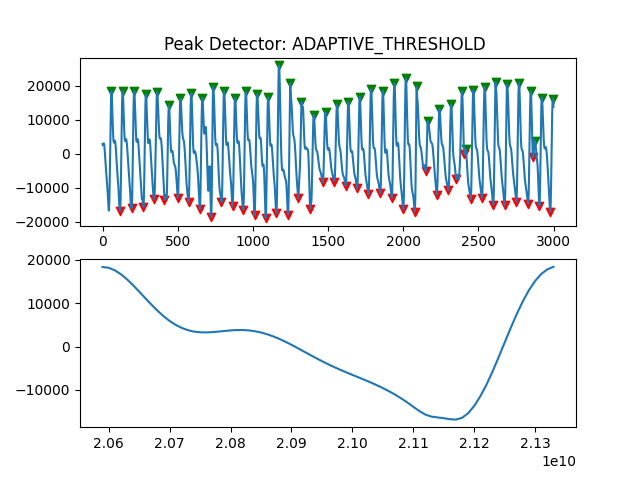
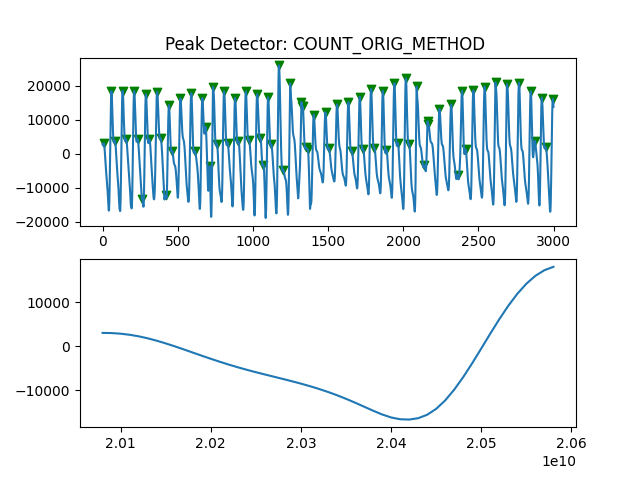
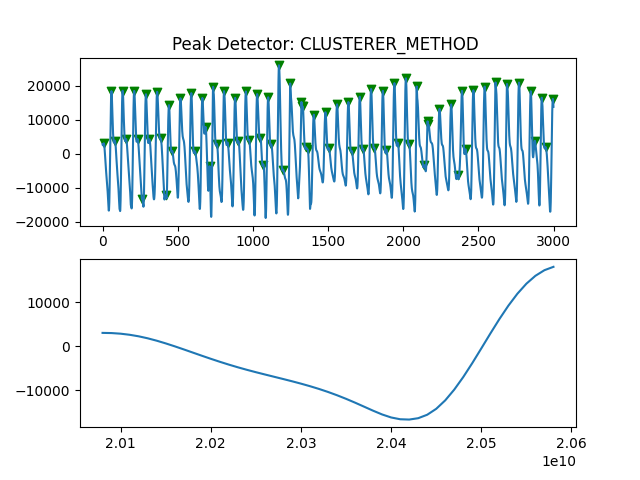
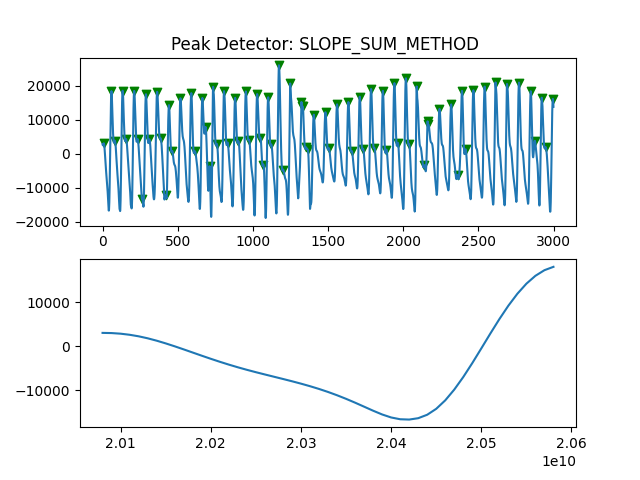
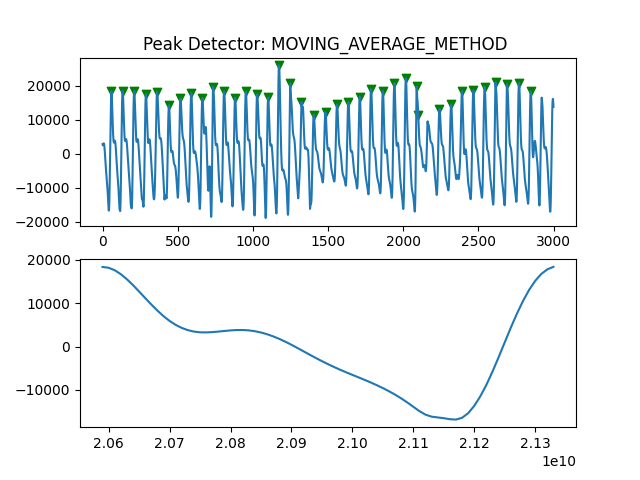
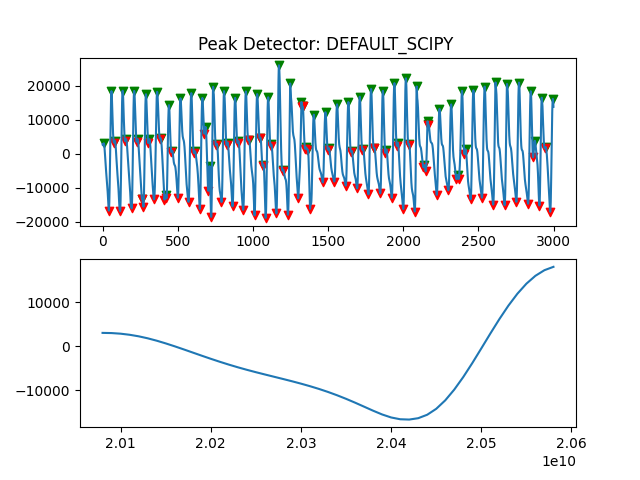
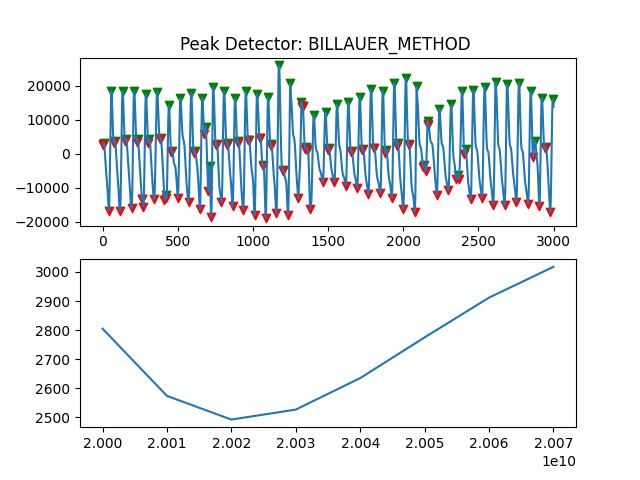
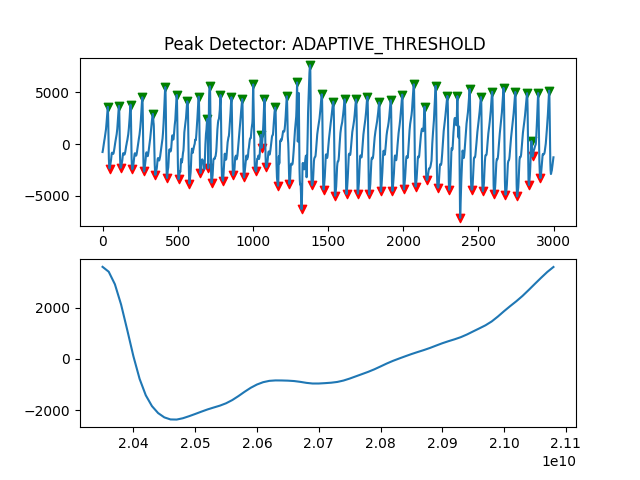
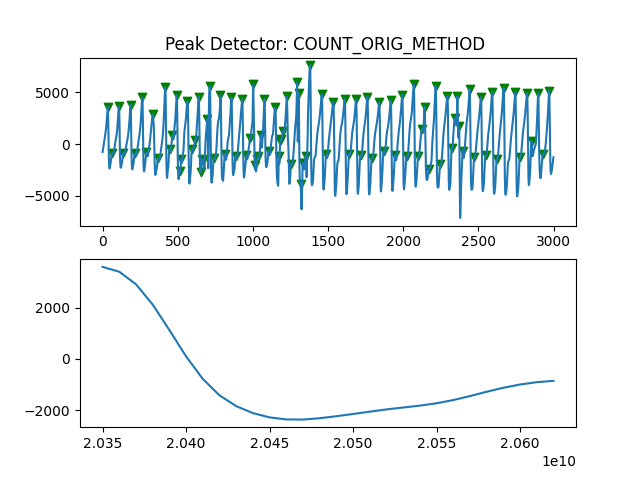
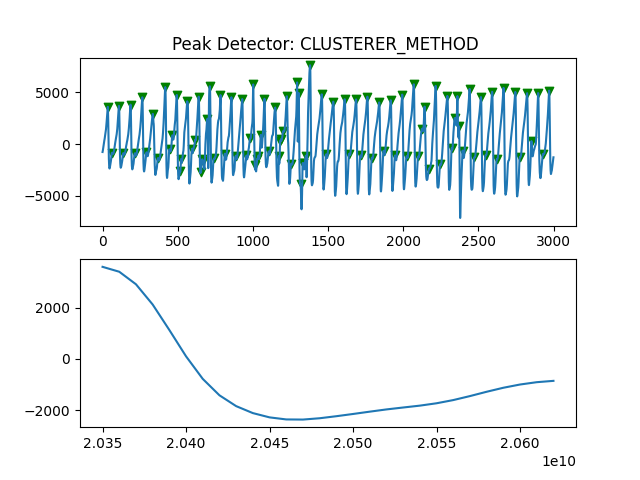
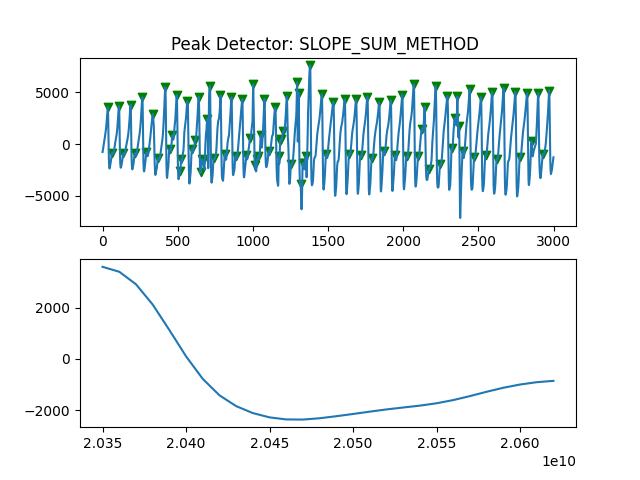
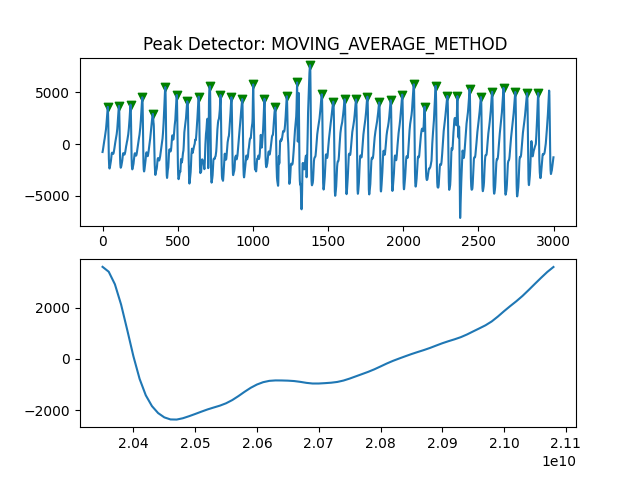
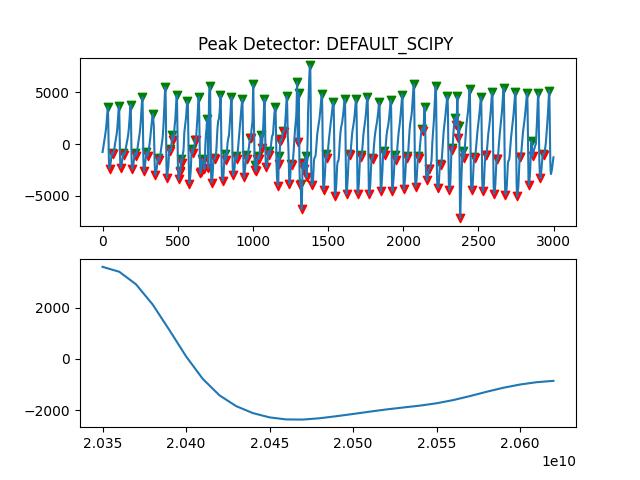
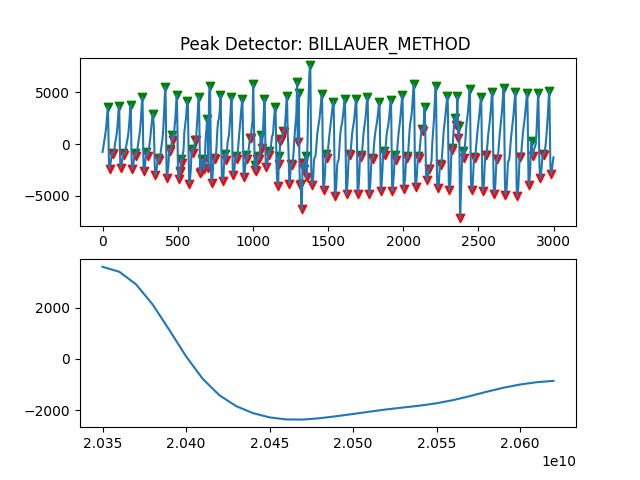
Peak Detector Comparison on PLETH
112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 | print('Peak Detector Comparison on Pleth')
pleth_segment= signals['Filtered_Pleth'][2000:5000]
for i in range(1,8,1):
peaks, troughs = PeakDetector().ppg_detector(pleth_segment, i)
sample_range = np.arange(0,3000,1)
fig, axes = plt.subplots(nrows=2, ncols=1)
axes = axes.flatten()
ax = axes[0]
ax.plot(sample_range, pleth_segment)
ax.scatter(peaks,pleth_segment[peaks], color="g", marker="v")
ax.scatter(troughs,pleth_segment[troughs], color="r", marker="v")
ax.title.set_text("Peak Detector: %s " %PeakDetectorNames[i-1])
ax = axes[1]
ax.plot(pleth_segment[peaks[0]:peaks[1]]) #plotting only one period
plt.show
|
Out:
Peak Detector Comparison on Pleth
the 'mode' parameter is not supported in the pandas implementation of take()
the 'mode' parameter is not supported in the pandas implementation of take()
search_for_onset() takes 3 positional arguments but 4 were given
<function show at 0x000001F0FFC0BD30>
Peak Detector Comparison on IR_ADC
134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 | print("Peak Detector Comparison on IR_ADC")
IRADC_segment= signals['Filtered_IR_ADC'][2000:5000]
for i in range(1,8,1):
peaks, troughs = PeakDetector().ppg_detector(IRADC_segment, i)
sample_range = np.arange(0,3000,1)
fig, axes = plt.subplots(nrows=2, ncols=1)
axes = axes.flatten()
ax = axes[0]
ax.plot(sample_range, IRADC_segment)
ax.scatter(peaks,IRADC_segment[peaks], color="g", marker="v")
ax.scatter(troughs,IRADC_segment[troughs], color="r", marker="v")
ax.title.set_text("Peak Detector: %s " %PeakDetectorNames[i-1])
ax = axes[1]
ax.plot(IRADC_segment[peaks[0]:peaks[1]]) #plotting only one period
plt.show
|
Out:
Peak Detector Comparison on IR_ADC
the 'mode' parameter is not supported in the pandas implementation of take()
the 'mode' parameter is not supported in the pandas implementation of take()
search_for_onset() takes 3 positional arguments but 4 were given
<function show at 0x000001F0FFC0BD30>
156 157 158 159 160 161 162 | '''
Detector 6 and 7 seem to be performing better overall
Note that the diacrotic notch can be higher than the diastolic peak
and therefore, cannot be effectively detected by the peak detector.
'''
|
Out:
'\n\nDetector 6 and 7 seem to be performing better overall\n\nNote that the diacrotic notch can be higher than the diastolic peak\nand therefore, cannot be effectively detected by the peak detector.\n'
Total running time of the script: ( 0 minutes 28.699 seconds)